Multiplex Immunohistochemical Phenotyping of T cells in Primary Prostate Cancer
Date: 20 October 2022
Time: 8:00 – 9:00 PST | 11:00 – 12:00 EST | 16:00 – 17:00 GMT
Location: Webinar
Learn about an iterative chromogenic based Serial Stain workflow in HALO®
Summary
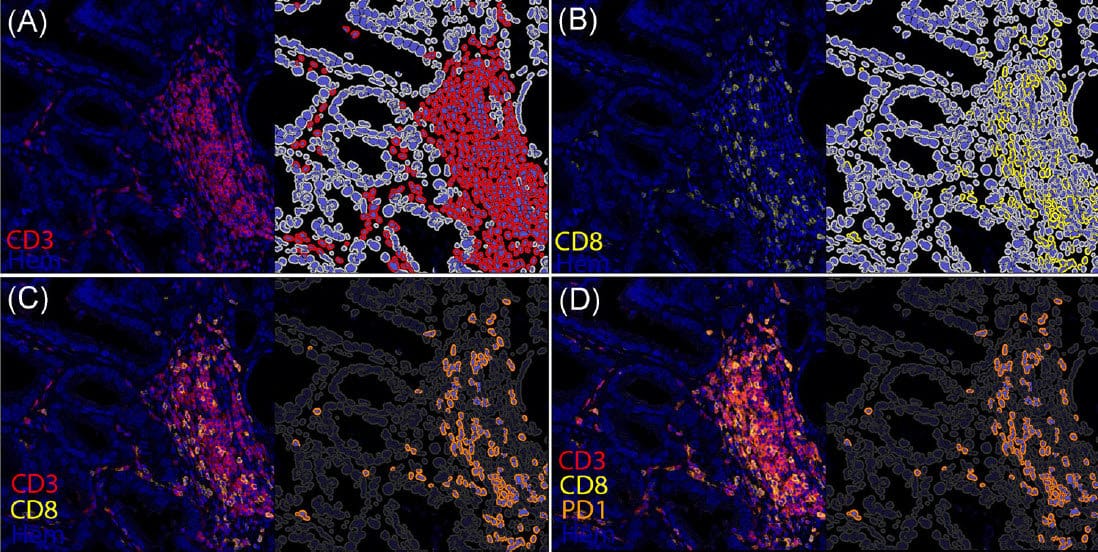
Multiplex immunohistochemistry is a powerful tool to visualize the spatial composition of inflammatory cells in the tumor microenvironment. Prostate cancer generally fits into the category of cancers that do not respond to immune checkpoint inhibitors. This may relate to the finding that most prostate cancers are known to be “immune cold” tumors. Nevertheless, the mechanisms leading to this immune desert-like state are not fully understood. We have designed a multiplex immunohistochemistry T cell panel to begin to systematically investigate the adaptive immune response in prostate cancer. We implemented an iterative chromogenic IHC assay approach, in which whole pathology slides are subjected to IHC and then scanned. Coverslips are removed and slides are then subjected to dissolution of the chromogen, antibody removal, second round antibody staining and rescanning. Iteratively stained whole slides scans are color deconvoluted and fused into a single multiplex image in HALO®. For analysis, we trained the Random Forest Tissue Classifier to define the stromal and epithelial components of the tumor separately, and used the HighplexFL module of HALO to quantify the densities of eight T cell phenotypes in the tumor epithelial and stromal subcompartments. Loss of PTEN is associated with the development of aggressive and metastatic prostate cancer. One of the mechanisms by which PTEN is postulated to drive disease aggressiveness is via immune suppression. Therefore, we used this panel to explore the association between the PTEN status of the tumor and the T cell densities in the tumor microenvironment, by comparing matched regions of tumors that showed PTEN genomic alterations and those with intact PTEN, within the tumor nodule.
Learning Objectives
- Learn about an iterative chromogenic based Serial Stain workflow in HALO
- Learn about validation of an iterative chromogenic assay in prostate cancer
Presenters

Busra Ozbeck, MD
Clinical Pathologist,
Derince Research and Training Hospital
Busra Ozbek is a Pathologist working in the University of Health Sciences Derince Research and Training Hospital, in Derince, Kocaeli, Turkey. She received her M.D. from Istanbul University. She completed her residency in pathology at the Kocaeli University, Faculty of Medicine. She did her postdoctoral training in the Johns Hopkins University School of Medicine, where she was the lead author on a paper that used whole slide images of iterative multiplex chromogenic IHC to study T cell infiltrates in prostate cancer (https://pubmed.ncbi.nlm.nih.gov/35188986/). She has experience in urological pathology, renal pathology, molecular pathology and computer-assisted image analysis methods. Her current research focuses on the multiplex immunohistochemistry and the role of inflammation in the pathogenesis and prognosis of prostate cancer.

Angelo M De Marzo MD, PHD
Professor of Pathology, Urology, and Oncology,
The Johns Hopkins University School of Medicine
Dr. Angelo De Marzo is a professor of pathology, oncology and urology at the Johns Hopkins University School of Medicine. He is board certified in Anatomic Pathology and for the past 20 years, his primary areas of clinical expertise are prostate pathology and molecular pathology. Dr. De Marzo serves as the Associate Director of Cancer Research Pathology at the Sidney Kimmel Comprehensive Cancer Center at Johns Hopkins.
Dr. De Marzo received his B.A. in molecular, cellular and developmental biology and psychology from the University of Colorado at Boulder. He earned his M.D. and Ph.D. in experimental pathology from University of Colorado Health Sciences Center. He completed his residency in pathology at the Johns Hopkins School of Medicine and performed a fellowship at the Brady Urological Institute at Johns Hopkins. Dr. De Marzo joined the Johns Hopkins faculty in 1998.
Dr. De Marzo’s research laboratory’s focus is on the molecular pathology of prostate cancer. He and colleagues described a potential new pathway for the molecular pathogenesis of prostate cancer, in which chronic inflammation results in epithelial cell damage and regeneration in regions termed proliferative inflammatory atrophy. Over time, this results in molecular alterations, including the emergence of luminal epithelial cells that overexpress c-MYC (MYC), that leads to the development of high grade prostatic intraepithelial neoplasia (PIN), or, at times directly to invasive carcinoma. His laboratory studies how MYC may drive the development of PIN and early carcinoma and has shown that MYC overexpression drives a key signature change consisting of increased nucleolar size and activity, and telomerase RNA (TERC) overexpression. Along with colleagues, including Dr. Srinvasan Yegnasubramanian, his lab studies transcriptomics and genomics of all stages of prostate cancer and precursor lesions. Working with mouse models and human tissues, he and collaborators are also exploring the role of chronic inflammation in driving prostatic cancer formation and progression and the synergy between MYC activation and PTEN loss in prostate cancer.
Dr. De Marzo’s lab also studies the prostate tumor microenvironment using multiplex IHC and in situ hybridization and his group has extensive experience validating IHC and in situ assays. The current work sheds light on the need to carefully annotate and exclude benign prostate tissue when studying the TME of prostate cancer.
Dr. De Marzo has published more than 370 papers, reviews and book chapters. He recently completed a 5 year term on the Editorial Board of the Journal of Clinical Investigation. He also serves on the Editorial Board for Cancer Prevention Research and The Prostate. He is a member of the Association of American Physicians. He served for several years as faculty at the AACR-ASCO Methods in Clinical Cancer Research Workshop, and currently serves on the AACR Science Education and Career Enhancement Committee, and the AACR Pathology Task Force.