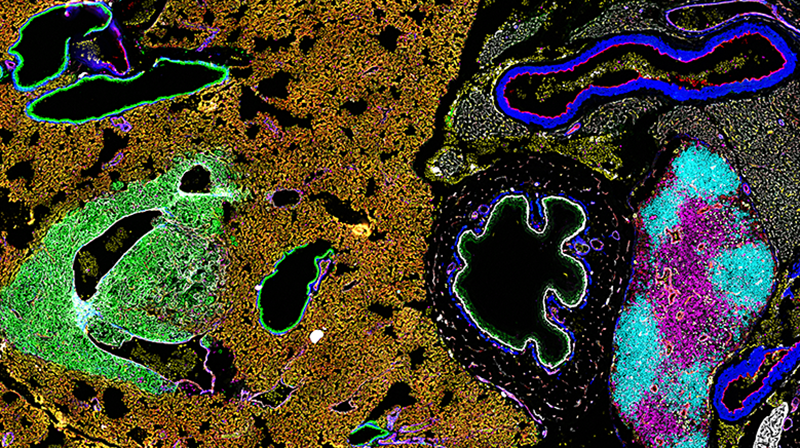
Optimizing quantitative analysis of highly multiplexed CODEX images in HALO
Date: 31 March 2022
Time: 8:00 – 9:00 PST | 11:00 – 12:00 EST | 16:00 – 17:00 GMT
Location: Webinar
CODEX technology (CO-Detection by indEXing) from Akoya Biosciences enables detection of up to 40 protein markers in one tissue with spatial context using DNA-conjugated antibodies and cyclic addition/removal of complementary fluorescently labeled DNA probes for visualization.
Summary
In this webinar, Dr. Noemi Kedei, a staff scientist at the Collaborative Protein Technology Resource core of the Center for Cancer Research (CCR) at the National Cancer Institute, will present a comprehensive CODEX workflow, from image acquisition to image processing and HALO image analysis. Dr. Kedei leads the core facility that provides this technology as service to CCR investigators, including project consultation and design, antibody panel customization, tissue staining and imaging, image processing, and data management. Since Dr. Kedei and team were an early adopter of the CODEX platform, they have acquired deep expertise in CODEX, having imaged hundreds of mouse and human fresh frozen and FFPE tissues in support of multiple NCI/CCR projects.
Dr. Kedei will present a technical webinar focused on the acquisition of CODEX images, image processing steps, and image analysis in HALO. She will discuss various HALO algorithms and modules such as the Highplex FL module and the Spatial Analysis modules, as well as the HALO AI Nuclei Segmenter and how they are used to generate quantitative single cell data and spatial analysis results.
Learning Objectives
- Acquire and process CODEX images for optimal performance in HALO®
- Generate quantitative single cell data and spatial analyses with HALO algorithms and modules, such as Highplex FL and Spatial Analysis
- Embed HALO AI™ nuclear segmentation into the Highplex FL module and combine with membrane markers to improve identification of single cells
Presenters

Noemi Kedei
Staff Scientist at the Collaborative Protein Technology Resource core,
NCI
Noemi is a staff scientist at the Collaborative Protein Technology Resource core at the CCR leading the core operations since 2020. The group is responsible for evaluating and implementing cutting edge immunoassays and innovative proteomic analysis technologies to facilitate basic discovery and translational research at CCR. With her key contributions, CODEX multiplex imaging as well as GeoMx Digital Spatial Profiling, two highly multiplex imaging platforms for detection of protein and RNA in tissues, are now offered as service to CCR investigators. The group continues to optimize all aspects of these technologies working with vendors to improve reagents, instrumentation, and software. Noemi is one of the NCI HALO Team leads and takes an active role in providing guidance and support to NCI users for digital image analysis.
Noemi got her MD degree from University of Debrecen, Hungary. She did her postdoctoral training and worked later as staff scientist in the Laboratory of Cancer Biology and Genetics, NCI. Her research focused on identifying mechanisms involved in the differential effect of natural compounds and their engineered derivatives acting on protein kinase Cs (PKCs), validated targets for anti-cancer treatment.