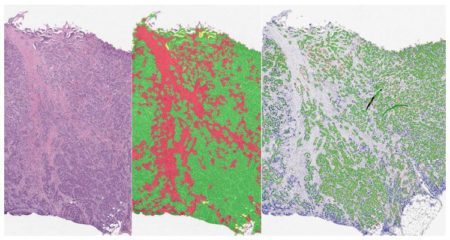
Leverage the Serial Section Analysis Add-on for HALO® to streamline your workflows for both serial section analysis and serial stain fusion analysis. For serial section analysis, the add-on enables users to create a tissue classifier based off a reference slide, such as an H&E image, and apply the resulting classification mask as regions of interest in serial section(s) stained for additional IHC markers. Serial section analysis is compatible with both brightfield and fluorescent images, and any of our HALO modules can be used for the final analysis step.
The serial stain fusion component of this add-on facilitates analysis of single slides that have been stained, stripped, and restained. In this workflow, brightfield images are deconvolved into their composite stains and converted to pseudo-fluorescent images before subsequent registration using a customizable set of configuration settings. Fluorescent images can also be registered using this workflow, just skip the deconvolution step. Lastly, the registered serial stain images can be fused into a single composite fluorescent image that can be analyzed using Highplex FL or any of our other fluorescent modules.
Want to know more? Contact us at info@indicalab.com to discuss your options with an applications scientist.

- Non-proprietary (JPG, TIF, OME.TIFF) (JPG, TIF, OME.TIFF)
- Nikon (ND2)
- 3D Histech (MRXS)
- Akoya Biosciences/Quanterix (QPTIFF, component TIFF)
- Olympus / Evident (VSI)
- Hamamatsu (NDPI, NDPIS)
- Aperio/Leica Biosystems (SVS, AFI)
- Zeiss (CZI)
- Leica (SCN, LIF)
- Ventana/Roche (BIF)
- Philips (iSyntax, i2Syntax)
- KFBIO (KFB, KFBF)
- DICOM (DCM*)
*whole-slide images

Sequential Same Slide mIF and H&E eBook
Reveal more data on a single slide with this workflow for sequential same slide multiplex IF and H&E staining and imaging using HALO and HALO AI.

mIF Co-Registration and Cellular Analysis eBook
Learn how HALO offers a streamlined workflow for co-registration and cellular analysis of immunofluorescence images from cyclically stained and imaged slides.

Qualitative and Quantitative Evaluation of the TME eBook
Check out this ebook to see how AI-based nuclear segmentation and tissue classification can streamline cell phenotyping across tissue compartments and samples.

CD38 in the Prostate Tumor Microenvironment
6 October 2021 | CD38, a druggable ectoenzyme, is involved in the generation of adenosine, which is implicated in tumor immune evasion. Its expression and

Webinar | Digital Pathology in the New Normal: Leveraging HALO® to Investigate a Global Pandemic
21 January 2021 | 8:00 AM – 9:00 AM PST | 11:00 AM – 12:00 PM EST | 4:00 PM – 5:00 PM GMT |<br

HALO® IMAGE ANALYSIS MASTERCLASS WEBINAR SERIES 2020
Summer of 2020 Indica Labs is excited to introduce our new HALO® Masterclass Webinar Series. Each masterclass webinar will offer a deep dive into a

The Use of Image Fusing in the Deployment of a 7-plex Immunofluorescent Assay
27 April 2020 | In lieu of our participation in the now cancelled 2020 AACR Conference, Indica Labs are pleased to bring you webinar presentations
Publication Spotlight
The table below includes publications that cite the Serial Section Analysis Add-on.
Your publication not on the list? Drop us an email to let us know about it!
| Title | Authors | Year | Journal | Topics | HALO Modules | Products |
|---|---|---|---|---|---|---|
| Neoadjuvant radioimmunotherapy in pancreatic cancer enhances effector T cell infiltration and shortens their distances to tumor cells | WANG J, GAI J, ZHANG T, NIU N, QI H, THOMAS II D L, LI K, XIA T, RODRIGUEZ C, PARKINSON R, DURHAM J, MCPHAUL T, NARANG A K, ANDERS R A, OSIPOV A, WANG H, HE J, LAHERU D A, HERMAN J M, LEE V, JAFFEE E M, THOMPSON E D, ZHU Q, ZHENG L | 2024 | Science Advances | Immuno-oncology | Spatial Analysis, Highplex FL, Registration | HALO |
| Tertiary lymphoid structures and B cells determine clinically relevant T cell phenotypes in ovarian cancer | Kasikova L, Rakova J, Hensler M, Lanickova T, Tomankova J, Pasulka J, Drozenova J, Mojzisova K, Fialova A, Vosahlikova S, Laco J, Ryska A, Dundr P, Kocian R, Brtnicky T, Skapa P, Capkova L, Kovar M, Prochazka J, Praznovec I, Koblizek V, Taskova A, Tanaka H, Lischke R, Mendez F C, Jr J V, Heinzelmann-Schwarz V, Jacob F, McNeish I A, Halaska M J, Rob L, Cibula D, Orsulic S, Galluzzi L, Spisek R, Fucikova J | 2024 | Nature Communications | Oncology | Classifier, Highplex FL, Registration | HALO |
| Development of a dual channel detection system for pan-genotypic simultaneous quantification of hepatitis B and delta viruses | Liu Y, Maya S, Carver S, O’Connell AK, Tseng AE, Gertje HP, Seneca K, Nahass RG | 2024 | Emerging Microbes & Infections | Infectious Disease | Registration | HALO |
| Pan-cancer single-cell dissection reveals phenotypically distinct B cell subtypes | Yang Y, Chen X, Pan J, Ning H, Zhang Y, Bo Y, Ren X, Li J, Qin S, Wang D, Chen M, Zhang Z | 2024 | Cell | Immuno-oncology | Spatial Analysis, Highplex FL, Registration | HALO |
| Chemotherapy drives tertiary lymphoid structures that correlate with ICI-responsive TCF1+CD8+ T cells in metastatic ovarian cancer | Lanickova T, Hensler M, Kasikova L, Vosahlikova S, Angelidou A, Pasulka J, Griebler H, Drozenova J, Mojzisova K, Vankerckhoven A, Laco J, Ryška A, Dundr P, Kocian R, Cibula D, Brtnicky T, Skapa P, Jacob F, Kovar M, Praznovec I, McNeish I A, Halaska M J, Rob L, Coosemans A, Orsulic S, Galluzzi L, Spisek R, Fucikova J | 2024 | Clinical Cancer Research | Immuno-oncology | Classifier, Highplex FL, Registration | HALO |
| Distinct immune cell infiltration patterns in pancreatic ductal adenocarcinoma (PDAC) exhibit divergent immune cell selection and immunosuppressive mechanisms | Sivakumar S, Jainarayanan A, Arbe-Barnes E, Sharma P K, Leathlobhair M N, Amin S, Reiss D J, Heij L, Hegde S, Magen A, Tucci F, Sun B, Wu S, Anand N M, Slawinski H, Revale S, Nassiri I, Webber J, Hoeltzel G D, Frampton A E, Wiltberger G, Neumann U, Charlton P, Spiers L, Elliott T, Wang M, Couto S, Lila T, Sivakumar P V, Ratushny A V, Middleton M R, Peppa D, Fairfax B, Merad M, Dustin M L, Abu-Shah E, Bashford-Rogers R | 2025 | Nature Communications | Immuno-oncology | Classifier, Multiplex IHC, Registration | HALO |
| Nanrilkefusp alfa (SOT101), an IL-15 receptor βγ superagonist, as a single agent or with anti-PD-1 in patients with advanced cancers | Champiat S, Garralda E, Galvao V, Cassier P A, Gomez-Roca C, Korakis I, Grell P, Naing A, LoRusso P, Mikyskova R, Podzimkova N, Reinis M, Ouali K, Schoenenberger A, Kiemle-Kallee J, Tillmanns S, Sachse R, Moebius U, Spisek R, Bechard D, Jelinkova L P, Adkins I, Marabelle A | 2025 | Cell Reports Medicine | Immuno-oncology | Spatial Analysis, Highplex FL, Registration | HALO |
| Spatial mapping of innate lymphoid cells in human lymphoid tissues and lymphoma at single-cell resolution | Van Acker N, Frenois F-X, Gravelle P, Tosolini M, Syrykh C, Laurent C, Brousset P | 2025 | Nature Communications | Immuno-oncology | Spatial Analysis, Highplex FL, Registration | HALO |
| Multidimensional, quantitative assessment of PD-1/PD-L1 expression in patients with Merkel cell carcinoma and association with response to pembrolizumab | Giraldo NA, Nguyen P, Engle EL, Kaunitz GJ, Cottrell TR, Berry S, Green B, Soni A, Cuda JD, Stein JE, Sunshine JC, Succaria F, Xu H, Ogurtsova A, Danilova L, Church CD, Miller NJ, Fling S, Lundgren L, Ramchurren N, Yearley JH, Lipson EJ, Cheever M, Anders RA, Nghiem PT, Topalian SL, Taube JM | 2018 | Journal for ImmunoTherapy of Cancer | Oncology, Immuno-oncology | Spatial Analysis, Serial Section | HALO |
| Empiric Methods to Account for Pre-analytical Variability in Digital Histopathology in Frontotemporal Lobar Degeneration | Giannini LAA, Xie SX, Peterson C, Zhou C, Lee EB, Wolk DA, Grossman M, Trojanowski JQ, McMillan CT, Irwin DJ | 2019 | Frontiers in Neuroscience | Neuroscience | Area Quantification, Registration, Figure Maker | HALO |
| Neoadjuvant immunotherapy leads to pathological responses in MMR-proficient and MMR-deficient early-stage colon cancers | Chalabi M, Fanchi LF, Dijkstra KK, Van den Berg JG, Aalbers AG, Sikorska K, Lopez-Yurda M, Grootscholten C, Beets GL, Snaebjornsson P, Maas M, Mertz J, Veninga V, Bounova G, Broeks A, Beets-Tan RG, de Wijkerslooth TR, van Lent AU, Marsman HA, Nuijten E, Kok NF, Kuiper M, Verbeek WH, Kok M, Van Leerdam ME, Schumacher TN, Voest EE, Haanen JB | 2020 | Nature Medicine | Immuno-oncology | Multiplex IHC, Serial Section | HALO |
| Assessment of associations between clinical and immune microenvironmental factors and tumor mutation burden in resected nonsmall cell lung cancer by applying machine learning to whole-slide images | Ono A, Terada Y, Kawata T, Serizawa M, Isaka M, Kawabata T, Imai T, Mori K, Muramatsu K, Hayashi I, Kenmotsu H, Oshima K, Urakami K, Nagashima T, Kusuhara M, Akiyama Y, Sugino T, Ohde Y, Yamaguchi K, Takahashi T T | 2020 | Cancer Medicine | Immuno-oncology | Classifier, Multiplex IHC, Serial Section | HALO |
| Single-cell analysis of germinal-center B cells informs on lymphoma cell of origin and outcome | Holmes AB, Corinaldesi C, Shen Q, Kumar R, Compagno N, Wang Z, Nitzan M, Grunstein E, Pasqualucci L, Dalla-Favera R, Basso K | 2020 | Journal of Experimental Medicine | Oncology | Registration | HALO |
| Multiplex Immunohistochemical Phenotyping of T Cells in Primary Prostate Cancer | Ozbek B, Ertunc O, Erickson A, Vidal I, Alexandre C, Guner G, Hicks J, Jones T, Taube J, Sfanos K, Yegnasubramanian S, DeMarzo A | 2021 | medRxiv | Immuno-oncology | Classifier, Cytonuclear, Highplex FL, Serial Section, Registration | HALO |
| SeqStain is an efficient method for multiplexed, spatialomic profiling of human and murine tissues | Rajagopalan A, Venkatesh I, Aslam R, Kirchenbuechler D, Khanna S, Cimbaluk D, Kordower JH, Gupta V | 2021 | Cell Reports Methods | Other | Spatial Analysis, Highplex FL, Registration | HALO |
| African-Ancestry Associated Gene Expression Signatures and Pathways in Triple Negative Breast Cancer, a Comparison across Women of African Descent | Martini R, Delpe P, Chu T, Arora K, Lord B, Verma A, Chen Y, Gebregzabher, Oppong J, Adjei E, Jibril A, Awuah B, Bekele M, Abebe E, Kyei I, Aitpillah F, Adinku M, Ankomah K, Osei-Bonsu E, Chitale D, Bensenhaver J, Nathanson S, Jackson L, Jiagge E, Petersen L, Proctor E, Gyan K, Gibbs L, Monojlovic Z, Kittles R, White J, Yates C, Manne U, Gardner K, Mongan N, Cheng E, Ginter P, Hoda S, Elemento O, Robine N, Sboner A, Carpten J, Newman L, Davis M | 2022 | medRxiv | Oncology | Multiplex IHC, Serial Section | HALO |
| Prostate cancer androgen receptor activity dictates efficacy of Bipolar Androgen Therapy | Sena L, Kumar R, Sanin D, Thompson E, Rosen D, Dalrymple S, Antony L, Yang Y, Comes-Alexandre C, Hicks J, Jones T, Bowers K, Eskra J, Meyers J, Gupta A, Skaist A, Yegnasubramanian S, Luo J, Brennen W, Kachhap S, Antonarakis E, De Marzo A, Isaacs J, Markowski M, Denmeade S | 2022 | medrxiv | Oncology | Classifier, Cytonuclear, Multiplex IHC, Registration | HALO |
| Metastatic recurrence in colorectal cancer arises from residual EMP1+ cells | Canellas-Socias A, Cortina C, Hernando-Momblona X, Palomo-Ponce S, Mulholland E, Turon G, Mateo L, Conti S, Roman O, Sevillano M, Slebe F, Stork D, Caballe-Mestres A, Berenguer-Llergo A, Alvarez-Varela A, Fenderico N, Novellasdemunt L, Jimenez-Gracia L, Sipka T, Bardia L, Lorden P, Colombelli J, Heyn H, Trepat X, Tejpar S, Sancho E, Tauriello D, Leedham S, Attolini C, Batlle E | 2022 | Nature | Oncology | Classifier, Highplex FL, Registration | HALO |
| Dynamic and adaptive cancer stem cell population admixture in colorectal neoplasia | Vasquez E, Nasreddin N, Valbuena G, Mulholland E, Belnoue-Davis H, Eggington H, Schenck R, Wouters V, Wirapati P, Gilroy K, Lannagan R, Flanagan D, Nujumudeen A, Omwenga S, McCorry A, Easton A, Koelzer V, East J, Morton D, Trusolino L, Maughan T, Campbell A, Loughrey M, Dunne P, Tsantoulis P, Juels D, Tejpar S, Sansom O, Leedham S | 2022 | Cell Stem Cell | Oncology | Registration | HALO |
| Tertiary lymphoid structures generate and propagate anti-tumor antibody-producing plasma cells in renal cell cancer | Meylan M, Petitprez F, Becht E, Bougouin A, Pupier G, Calvez A, Giglioli I, Verkarre V, Lacroix G, Verneau J, Sun C, Laurent-Puig P, Vano Y, Reynaud C, Reynies A, Sautes-Fridman C, Fridman W | 2022 | Immunity | Oncology | Spatial Analysis, Registration | HALO, HALO AI |
| A lncRNA signature associated with tumor immune heterogeneity predicts distant metastasis in locoregionally advanced nasopharyngeal carcinoma | Liang Y, Zhang Y, Tan X, Qiao H, Liu S, Tang L, Mao Y, Chen L, Li W, Zhou G, Zhao Y, Li J, Li Q, Huang S, Gong S, Zheng Z, Li Z, Sun Y, Jiang W, Ma J, Li Y, Liu N | 2022 | Nature Communications | Immuno-oncology | Multiplex IHC, Registration | HALO |
| Systemic Nos2 Depletion and Cox inhibition limits TNBC disease progression and alters lymphoid cell spatial orientation and density | Somasundaram V, Ridnour L, Cheng R, Walke A, Kedei N, Bhattacharyya D, Wink A, Edmondson E, Butcher D, Warner A, Dorsey T, Scheiblin D, Heinz W, Bryant R, Kinders R, Lipkowitz S, Wong S, Pore M, Hewitt S, McVicar D, Anderson S, Chang J, Glynn S, Ambs S, Lockett S, Wink D | 2022 | Redox Biology | Oncology | Classifier, Registration | HALO |
| PERK reprograms hematopoietic progenitor cells to direct tumor-promoting myelopoiesis in the spleen | Liu M, Wu C, Luo S, Hua Q, Chen H, Weng Y, Xu J, Lin H, Wang L, Li J, Zhu L, Guo Z, Zhuang S, Kang T, Zheng L | 2022 | Journal of Experimental Medicine | Oncology | Highplex FL, Registration | HALO |
| Assessment of spatial transcriptomics for oncology discovery | Lyubetskaya A, Rabe B, Fisher A, Lewin A, Neuhaus I, Brett C, Brett T, Pereira E, Golhar R, Kebede S, Font-Tello A, Mosure K, Wittenberghe N, Mavrakis K, Maclsaac K, Chen B, Drokhlyansky E | 2022 | Cell Reports Methods | Oncology | Registration | HALO, HALO AI |
| Unchecked oxidative stress in skeletal muscle prevents outgrowth of disseminated tumour cells | Crist S, Nemkov T, Dumpit R, Dai J, Tapscott S, True L, Swarbrick A, Sullivan L, Nelson P, Hansen K, Ghajar C | 2022 | Nature Cell Biology | Myology | Registration | HALO |
| Immune cells in mesothelioma microenvironment simplistic marker of response to nivolumab plus ipilimumab | Disselhorst M, Lubeck Y, van der Noort V, Quispel-Janssen J, Seignette I, Sanders J, Peters D, Hooijberg E, Baas P | 2022 | Lung Cancer | Oncology | Multiplex IHC, Registration | HALO |
| Multiplex Immunohistochemical Phenotyping of T Cells in Primary Prostate Cancer | Ozbek B, Ertunc O, Erickson A, Vidal I, Gomes-Alexandre C, Guner G, Hicks J, Jones T, Taube J, Sfanos K, Yegnasubramanian S, DeMarzo A | 2022 | The Prostate | Immuno-oncology | Classifier, Serial Section, Registration, Figure Maker | HALO |
| CXCL9 and CXCL13 coordinately shape the immune-activated microenvironment of endometrial cancer via tertiary lymphoid structure formation | Kodama M, Nagase Y, Aimono E, Nakamura K, Takamatsu R, Abe K, Yoshimura T, Chiyoda T, Yamagami W, Aoki D, Nishihara H | 2023 | Research Square | Immuno-oncology | Registration | HALO |
| MYC-driven increases in mitochondrial DNA copy number occur early and persist throughout prostatic cancer progression | Chen J, Zheng Q, Hicks J, Trabzonlu L, Ozbek B, Jones T, Vaghasia A, Larman T, Wang R, Markowski M, Denmeade S, Pienta K, Hruban R, Antonaraskis E, Gupta A, Dang C, Yegnasubramanian S, De Marzo A | 2023 | bioRxiv | Oncology | Classifier, Area Quantification, Serial Section | HALO |
| Fucosylation of HLA-DRB1 regulates CD4+ T cell-mediated anti-melanoma immunity and enhances immunotherapy efficacy | Lester D, Burton C, Gardner A, Innamarato P, Kodumudi K, Liu Q, Adhikari E, Ming Q, Williamson D, Frederick D, Sharova T, White M, Markowitz J, Cao B, Nguyen J, Johnson J, Beatty M, Mockabee-Macias A, Mercurio M, Watson G, Chen P-L, McCarthy S, MoranSegura C, Messina J, Thomas K, Darville L, Izumi V, Koomen J, Pilon-Thomas S, Ruffell B, Luca V, Haltiwanger R, Wang X, Wargo J, Boland G, Lau E | 2023 | Nature Cancer | Immuno-oncology | Registration | HALO |
| Inhibition of the proline metabolism rate-limiting enzyme P5CS allows proliferation of glutamine-restricted cancer cells | Linder S, Bernasocchi T, Martínez-Pastor B, Sullivan K, Galbraith M, Lewis C, Ferrer C, Boon R, Silveira G, Cho H, Vidoudez C, Shroff S, Oliveira-Costa J, Ross K, Massri R, Matoba Y, Kim E, Rueda B, Stott S, Gottlieb E, Espinosa J, Mostoslavsky R | 2023 | Nature Metabolism | Oncology, Metabolism | Multiplex IHC, Registration | HALO |
| Spatial Mapping of Human Hematopoiesis at Single Cell Resolution Reveals Aging-Associated Topographic Remodeling | Sarachakov A, Varlamova A, Svelolkin V, Polyakova M, Valencia I, Unkenholz C, Pannellini T, Galkin I, Ovcharov P, Tabakov D, Postovalova E, Shin N, Sethi I, Bagaev A, Itkin T, Crane GM, Kluk MJ, Geyer JT, Inghirami GGA, Patel SS | 2023 | Blood | Other | Registration | HALO |
| Quantitative intravital imaging for real-time monitoring of pancreatic tumor cell hypoxia and stroma in an orthotopic mouse model | Samuel T, Rapic S, O’Brien C, Edson M, Zhong Y, DaCosta R | 2023 | Science Advances | Oncology | Registration | HALO |
| A Cdh3-β-catenin-laminin signaling axis in a subset of breast tumor leader cells control leader cell polarization and directional collective migration | Hwang P, Mathur J, Cao Y, Almeida J, Ye J, Morikis V, Cornish D, Clarke M, Stewart S, Pathak A, Longmore G | 2023 | Developmental Cell | Oncology | Highplex FL, Registration | HALO |
| The localization of molecularly distinct microglia populations to Alzheimer’s disease pathologies using QUIVER | Shahidehpour R, Nelson A, Sanders L, Embry C, Nelson P, Bachstetter A | 2023 | Acta Neuropathologica Communications | Neuroscience | Area Quantification, Object Colocalization, Spatial Analysis, Registration | HALO |
| PD-1+CD8+ T Cells Proximal to PD-L1+CD68+ Macrophages Are Associated with Poor Prognosis in Pancreatic Ductal Adenocarcinoma Patients | Yang X, Wang G, Song Y, Zhuang T, Li Y, Xie Y, Fei X, Zhao Y, Xu D, Hu Y | 2023 | Cancers | Oncology, Immuno-oncology | Classifier, Spatial Analysis, Highplex FL, Registration | HALO |
| Gastric tubular adenocarcinoma with diffuse neutrophils infiltrating: characteristics and probable treatment strategy | Wang B, Zhu Y, Wang S, Li Z, Wang L, Rao W, Cheng N, Chen R, Ying J, Xue L | 2023 | Gastric Cancer | Oncology | Multiplex IHC, Spatial Analysis, Registration | HALO |
| Standardized pathology screening of mature Tertiary Lymphoid Structures in cancers | Vanhersecke L, Bougouin A, Crombe A, Brunet M, Sofeu C, Parrens M, Pierron H, Bonhomme B, Lembege N, Rey C, Velasco V, Soubeyran I, Begueret H, Bessede A, Bellera C, Scoazec J-Y, Italiano A, Fridman C, Fridman W, Le Loarer F | 2023 | Laboratory Investigation | Other | Highplex FL, Registration | HALO |
| Whole-Slide Imaging, Mutual Information Registration for Multiplex Immunohistochemistry and Immunofluorescence | Doyle J, Green B, Eminizer M, Jimenez-Sanchez D, Lu S, Engle E, Xu H, Ogurtsova A, Lai J, Soto-Diaz S, Roskes J, Deutsch J, Taube J, Sunshine J, Szalay A | 2023 | Laboratory Investigation | Other | Multiplex IHC, Registration | HALO |
| A machine learning approach toward automating spatial identification of LAG3+/CD3+ cells in ulcerative colitis | Bonnevie ED, Dobrzynski E, Steiner D, Hildebrand D, Monslow J, Singh M, Decman V, Krull DL | 2023 | Scientific Reports | Gastroenterology | Highplex FL, Registration | HALO, HALO AI |
| Cell signaling activation and extracellular matrix remodeling underpin glioma tumor microenvironment heterogeneity and organization | Dinevska M, Widodo S, Furst L, Cuzcano L, Fang Y, Mangiola S, Neeson P, Darcy P, Ramsay R, Hutchinson R, MacKay F, Christie M, Stylli S, Mantamadiotis T | 2022 | Cellular Oncology | Oncology | Classifier, Highplex FL, Registration | HALO |
Related HALO Modules
Quantify expression of up to five brightfield stains in any cellular compartment - membrane, nucleus or cytoplasm.
Learn MorePlot cells and objects from one or more images and perform nearest neighbor analysis, proximity analysis, and tumor infiltration analysis.
Learn MoreSeparate multiple tissue classes across a tissue using a learn-by-example approach. Can be used in conjunction with all other modules (fluorescent and brightfield) to select specific tissue classes for further analysis.
Learn MoreWant to Learn More?
Fill out the form below to request information about any of our software products.
You can also drop us an email at info@indicalab.com
Products & Services
Interested in purchasing or learning more about our products and services? Our highly trained application scientists are a couple of clicks away.
Software Maintenance & Support Coverage
Interested in purchasing an SMS plan? We would be happy to give you a quote.
Technical Support
Need technical support? Our IT specialists are here to help.