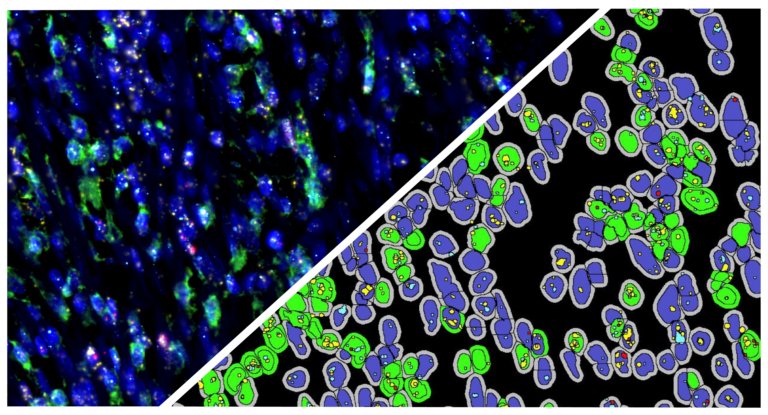
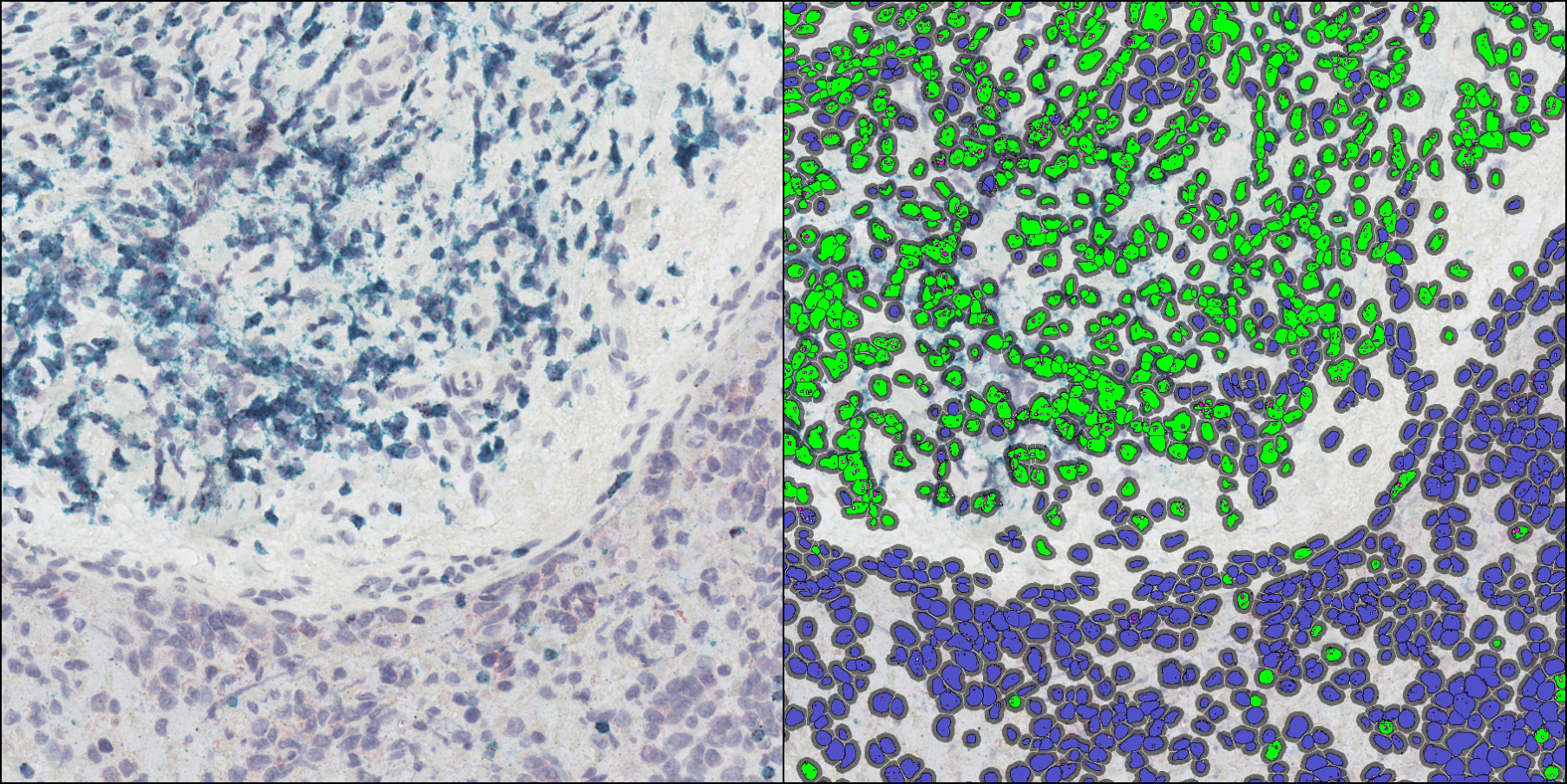
Use the HALO® ISH-IHC module and reagents from Molecular Instruments or ACD, a Bio-techne brand, to simultaneously analyze a nuclear stain and up to four IHC biomarkers or ISH probes on a cell-by-cell basis. This module enables in-depth analysis of the corresponding protein and gene expression profile of every cell across a brightfield image, reporting outputs including IHC positive cells in total and by cell compartment, probe copies and area per cell, and calculated H-scores for each probe. Optimize your image analysis with available pre-trained AI-based membrane and/or nuclear segmentation, now included with the HALO platform, and leverage interactive markups to dynamically explore colocalization and combined cell phenotypes.
Try it out! Click here to initiate your free proof-of-concept HALO image analysis.
Feature image provided by Molecular Instruments
Sample: Mouse Pancreas
Targets: Polr2a (RNA), HCR™ Membrane Stain (Protein)
Magnification: 20x
- Non-proprietary (JPG, TIF, OME.TIFF) (JPG, TIF, OME.TIFF)
- Nikon (ND2)
- 3D Histech (MRXS)
- Akoya Biosciences/Quanterix (QPTIFF, component TIFF)
- Olympus / Evident (VSI)
- Hamamatsu (NDPI, NDPIS)
- Aperio/Leica Biosystems (SVS, AFI)
- Zeiss (CZI)
- Leica (SCN, LIF)
- Ventana/Roche (BIF)
- Philips (iSyntax, i2Syntax)
- KFBIO (KFB, KFBF)
- DICOM (DCM*)
*whole-slide images

RNAscope™ and HALO® Publications
Read this publications list for examples of research in areas from neuroscience to oncology using HALO to analyze RNAscope™ images.

Quantitative RNAscope™ Image Analysis Guide
From experimental design considerations to optimized setup of HALO image analysis parameters, our guide will help take your quantitative RNAscope™ image analysis to the next level.

HCR™ Pro RNA-ISH Image Analysis Using HALO
Download our collaborative poster with Molecular Instruments to learn how membrane image analysis improved RNA-ISH quantification.

Getting Started with RNAscope™ Image Analysis in HALO®
28 March 2023 | Join us for this 1-hour webinar to see a live demonstration of RNAscope image analysis using the HALO® platform from Indica
Masterclass Webinar: Optimizing RNAscope Image Analysis
05 May 2022 | In this 60-min webinar, Dr. Ghislaine Lioux will present solutions to common RNAscope image analysis challenges including how to optimize color

HALO Image Analysis Masterclass Series: February-March 2021
Spring of 2021 | Indica Labs is excited to continue our HALO® Masterclass Webinar Series this winter. Each masterclass webinar will offer a deep dive

Webinar | Digital Pathology in the New Normal: Leveraging HALO® to Investigate a Global Pandemic
21 January 2021 | 8:00 AM – 9:00 AM PST | 11:00 AM – 12:00 PM EST | 4:00 PM – 5:00 PM GMT |<br
Publication Spotlight
The table below includes publications that cite the ISH-IHC and module.
Your publication not on the list? Drop us an email to let us know about it!
| Title | Authors | Year | Journal | Topics | HALO Modules | Products |
|---|---|---|---|---|---|---|
| Phase I trial of ADP-A2AFP TCR T-cell therapy in patients with advanced hepatocellular or gastric hepatoid carcinoma | Meyer T, Finn R S, Borad M, Mahipal A, Edeline J, Houot R, Hausner P F, Hollebecque A, Goyal L, Frigault M, Evans T R J, Wong K M, Tan B R, Mitry E, Sarker D, Feun L, El-Rayes B, Thistlethwaite F, Kaseb A, Alese O, Jin Z, Cirillo C, Bruix J, Roddie C, Noto P, Fayngerts S, Cristiani S, Sampson J, Bai J, Isabelle M, Broad R, Sun A, Norry E, Sangro B | 2025 | Journal of Hepatology | Immuno-oncology | Highplex FL, ISH-IHC | HALO |
| Detection of engineered T cells in FFPE tissue by multiplex in situ hybridization and immunohistochemistry | Wright JH, Huang L-Y, Weaver S, Archila LD, McAfee MS, Hirayama AV, Chapuis AG, Bleakley M, Rongvaux A, Turtle CJ, Chanthaphavong S, Campbell JS, Pierce RH | 2020 | Journal of Immunological Methods | Immunology | ISH/FISH, ISH-IHC/FISH-IF | HALO |
| Spatially organized multicellular immune hubs in human colorectal cancer | Pelka K, Hofree M, Chen J, Sarkizova S, Pirl J, Jorgji V, Bejnood A, Dionne D, Ge W, Xu K, Chao S, Zollinger D, Lieb D, Reeves J, Fuhrman C, Hoang M, Delorey T, Nguyen L, Waldman J, Klapholz M, Wakiro I, Cohen O, Albers, J, Smillie C, Cuoco M, Wu J, Su M, Yeung J, Vijaykumar B, Magnuson A, Asinovski N, Moll T, Goder-Reise M, Applebaum A, Brais L, DelloStritto L, Denning S, Phillips S, Hill E, Meehan J, Frederick D, Sharova T, Kanodia A, Todres E, Jane-Valbuena J, Biton M, Izar B, Lambden C, Clancy T, Bleday R, Melnitchouk N, Irani J, Kunitake H, Berger D, Srivastava A, Hornick J, Ogino S, Rotem A, Vigneau S, Johnson B, Corcoran R, Sharpe A, Kuchroo V, Ng K, Giannakis M, Nieman L, Boland G, Arguirre A, Anderson A, Rosenblatt-Rosen O, Regev A, Hachohen N | 2021 | Cell | Oncology | Spatial Analysis, ISH-IHC/FISH-IF | HALO |
| Detection of Epstein–Barr Virus in Periodontitis: A Review of Methodological Approaches | Tonoyan L, Chevalier M, Vincent-Bungnas S, Marsault R, Doglio A | 2021 | Mircroorganisms | Review | ISH/FISH, ISH-IHC/FISH-IF | HALO |
| Multiparameter immunohistochemistry analysis of HIV DNA, RNA and immune checkpoints in lymph node tissue | Richardson Z, Deleage C, Tutuka C, Walkiewica M, Del Rio-Estrada P, Pascoe R, Evans V, Reyesteran G, Gonzales M, Roberts-Thomson S, Gonzalez-Navarro M, Torres-Ruiz, F, Estes J, Lewin S, Cameron P | 2021 | Journal of Immunological Methods | Infectious Disease | ISH-IHC/FISH-IF | HALO |
| IL-1β Promotes Expansion of IL-33+ Lung Epithelial Stem Cells Following RSV Infection During Infancy | Vu L, Phan A, Hijano D, Siefker D, Tillman H, Cormierr S | 2021 | American Journal of Respiratory Cell and Molecular Biology | Infectious Disease | ISH-IHC | HALO |
| Satellite repeat RNA expression in epithelial ovarian cancer associates with a tumor immunosuppressive phenotype | Porter R, Sun S, Flores M, Berzolla E, You E, Phillips I, KC N, Desai N, Tai E, Szabolcs A, Lang E, Pankaj A, Raabe M, Thapar V, Xu K, Nieman L, Rabe D, Kolin D, Stover E, Pepin D, Stott S, Deshpande V, Liu J, Solovyov A, Matulonis U, Greenbaum B, Ting D | 2022 | The Journal of Clinical Investigation | Immuno-oncology | ISH/FISH, ISH-IHC/FISH-IF | HALO |
| Annexin A2/TLR2/MYD88 pathway induces arginase 1 expression in tumor-associated neutrophils | Zhang H, Zhu X, Friesen T, Kwak J, Pisarenko T, Mekvanich S, Velasco M, Randolph T, Kargl J, Houghton A | 2022 | Journal of Clinical Investigation | Immuno-oncology | ISH-IHC/FISH-IF | HALO |
| Spatiotemporal transcriptome analysis reveals critical roles for mechano-sensing genes at the border zone in remodeling after myocardial infarction | Yamada S, Ko T, Hatsuse S, Nomura S, Zhang B, Dai Z, Inoue S, Kubota M, Sawami K, Yamada T, Sassa T, Katagiri M, Fujita K, Katoh M, Ito M, Harada M, Toko H, Takeda N, Morita H, Aburatani H, Komuro I | 2022 | Nature Cardiovascular Research | Myology | ISH-IHC/FISH-IF | HALO |
| Loss of functional System x-c uncouples aberrant postnatal neurogenesis from epileptogenesis in the hippocampus of Kcna1-KO mice | Aloi M, Thompson S, Quartapella N, Noebels J | 2022 | Cell Reports | Neuroscience | ISH-IHC/FISH-IF | HALO |
| The potassium channel auxiliary subunit Kvβ2 (Kcnab2) regulates Kv1 channels and dopamine neuron firing | Yee J, Rastani A, Soden M | 2022 | Journal of Neurophysiology | Neuroscience | ISH-IHC/FISH-IF | HALO |
| Transcriptional vulnerabilities of striatal neurons in human and rodent models of Huntington’s disease | Matsushima A, Pineda S, Crittenden J, Lee H, Galani K, Mantero J, Tombaugh G, Kellis M, Heiman M, Graybiel A | 2023 | Nature Communications | Neuroscience | Area Quantification, ISH-IHC/FISH-IF | HALO |
Related HALO Modules
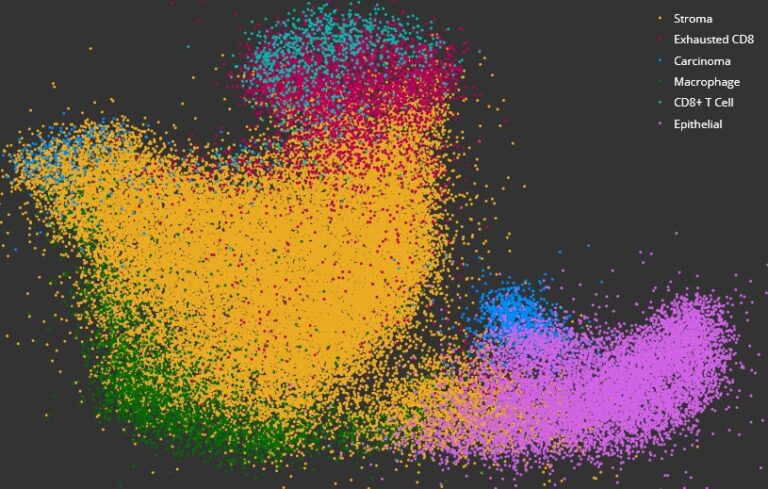
Acquire deeper insights into complex data sets using dimensional reduction and unsupervised clustering with interactive plotting.
Learn MoreUse the HALO® FISH-IF module and reagents from Molecular Instruments or ACD, a Bio-techne brand, to simultaneously analyze an unlimited number of fluorescently-labeled DNA/RNA ISH probes and immunofluorescent protein biomarkers on a cell-by-cell basis.
Learn MoreSimultaneously analyze up to three chromogenic and/or silver-labelled DNA or RNA ISH probes on a cell-by-cell basis, measuring spot numbers and area per cell and compartment, and calculated H-scores for each probe.
Learn MoreWant to Learn More?
Fill out the form below to request information about any of our software products.
You can also drop us an email at info@indicalab.com
Products & Services
Interested in purchasing or learning more about our products and services? Our highly trained application scientists are a couple of clicks away.
Software Maintenance & Support Coverage
Interested in purchasing an SMS plan? We would be happy to give you a quote.
Technical Support
Need technical support? Our IT specialists are here to help.